The Environmental Afterlife of Antibiotics
When we allow antibiotics to linger in the environment, we are maintaining a breeding ground for antibiotic resistance.

Read Time: 4 minutes
Published:
For most of human history, bacterial diseases like pneumonia or tuberculosis, and even infected wounds, were a death sentence. All that changed in the 1940s, when penicillin, the first drug that could treat bacterial infections, became widely available. Antibiotics were a miracle, and doctors, who had previously been unable to do much for these patients, prescribed them liberally.
Since then, antibiotic overuse (and misuse) has led some bacteria to develop the ability to survive the antibiotics designed to kill them, turning once-treatable infections back into hard-to-control threats. Today, these antibiotic-resistant bacteria directly cause more than 1 million deaths annually, and are projected to cause another 40 million more deaths between 2025 and 2050. The majority of these cases are expected to be driven by multidrug-resistant tuberculosis as well as the “big three”: E. coli, Staphylococcus aureus (MRSA), and bacterial pneumonia.
At the center of this crisis are antibiotic resistance genes, pieces of genetic code that allow bacteria to deactivate or excrete antibiotic drugs. Resistance genes thrive in hospitals, where antibiotics are constantly used to lower the risk of hospital-acquired infections. They are widespread in the agricultural industry, where they are used to preempt the spread of disease among livestock. And they are also amplified by exposure to environmental antibiotics.
Up to 90% of antibiotics are eventually excreted, unmetabolized, into the environment through wastewater and manure, making their way into soil and surface water. This means that bacteria in these environments are constantly exposed to low levels of antibiotics. In an environment with no antibiotics, bacteria would have no reason to develop or maintain resistance genes. However, in the presence of antibiotics, these genes are a survival advantage. Antibiotics kill off or weaken vulnerable bacteria, allowing those with resistance genes to survive and reproduce.
This risk exists because the different bacteria that affect humans, animals, and the environment do not stay in separate siloes.
In a new study, Yue Wang and colleagues explored how exposure to four common antibiotics (tetracycline, ampicillin, kanamycin, and streptomycin) affects the spread of antibiotic resistance genes in environmental bacteria. The researchers examined two methods of gene transfer: vertical transfer, when resistance is passed down from parent to offspring through reproduction, and horizontal transfer, when genes are swapped between unrelated bacteria. Gene swapping occurs either through direct contact between cells or when bacteria pick up loose DNA from their surroundings.
For 10 days, the researchers exposed one group of bacteria to low levels of antibiotics to simulate environmental concentrations, and another group to a typical clinical dose of antibiotics. They then used the data to model how antibiotic resistance would build up in the environment over longer time periods.
In their experiments, the clinical dose of antibiotics killed the bacteria, but low environmental levels did not. Instead, these low doses allowed resistant bacteria to survive and accumulate over time. Exposure to trace amounts of antibiotics also increased the likelihood that bacteria would retain resistance genes across generations (vertical transfer) and share them more efficiently with neighboring bacteria (horizontal transfer). Together, these results suggest that trace amounts of antibiotics in the environment contribute to the persistence and spread of resistance genes.
When we allow antibiotics to linger in the environment, we are maintaining a breeding ground for antibiotic resistance.
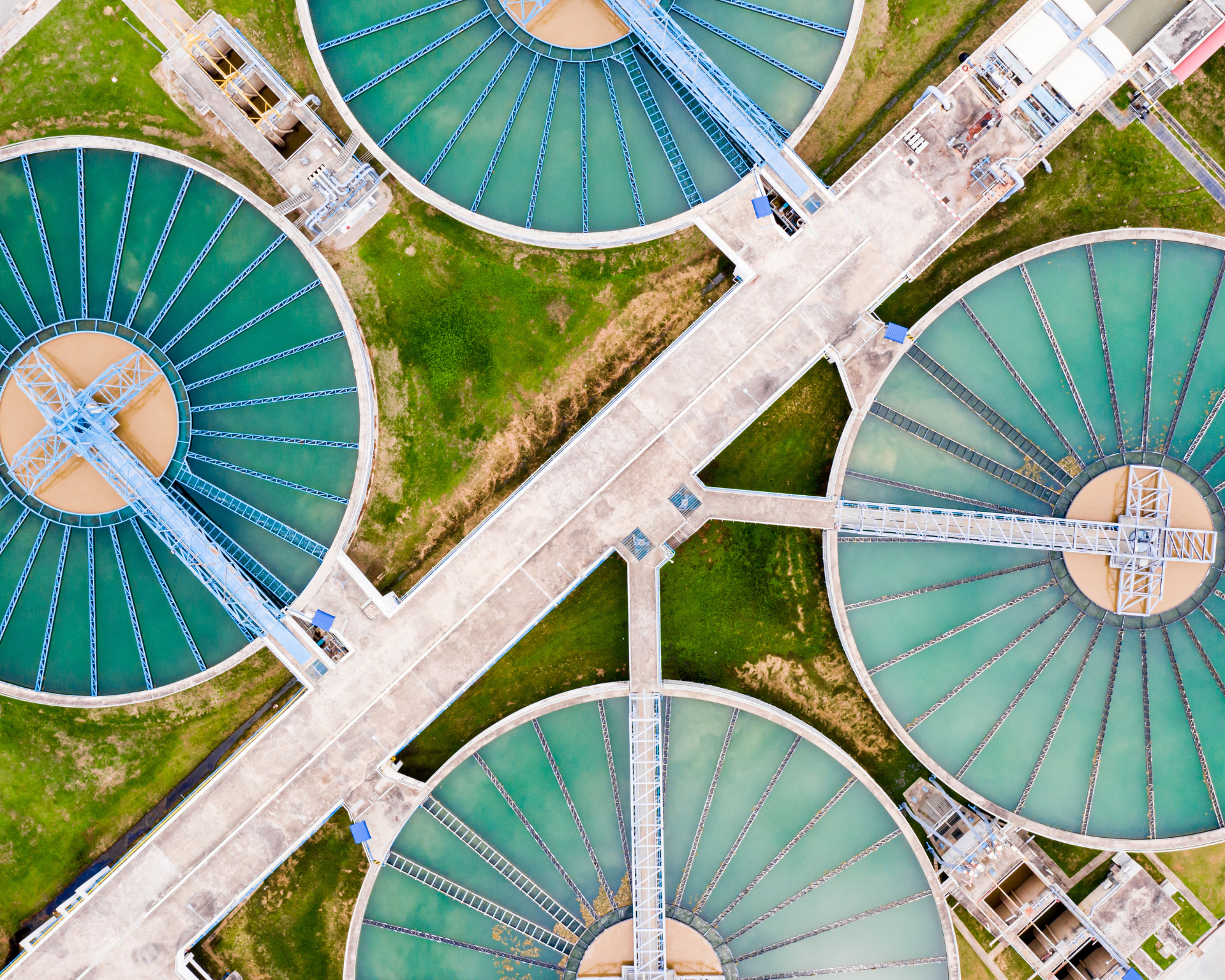
Addressing antibiotic overuse in medicine and agriculture alone will not solve antibiotic resistance. The researchers argue that antibiotics should be regulated as environmental contaminants, like pesticides or microplastics. While this study was conducted in a lab, its findings align with field data showing that places where environmental and disease-causing bacteria meet, like wastewater treatment plants, are hotspots for antibiotic-resistant infections. Previous research has also proposed environmental discharge thresholds and supported calls to regulate antibiotics as pollutants.
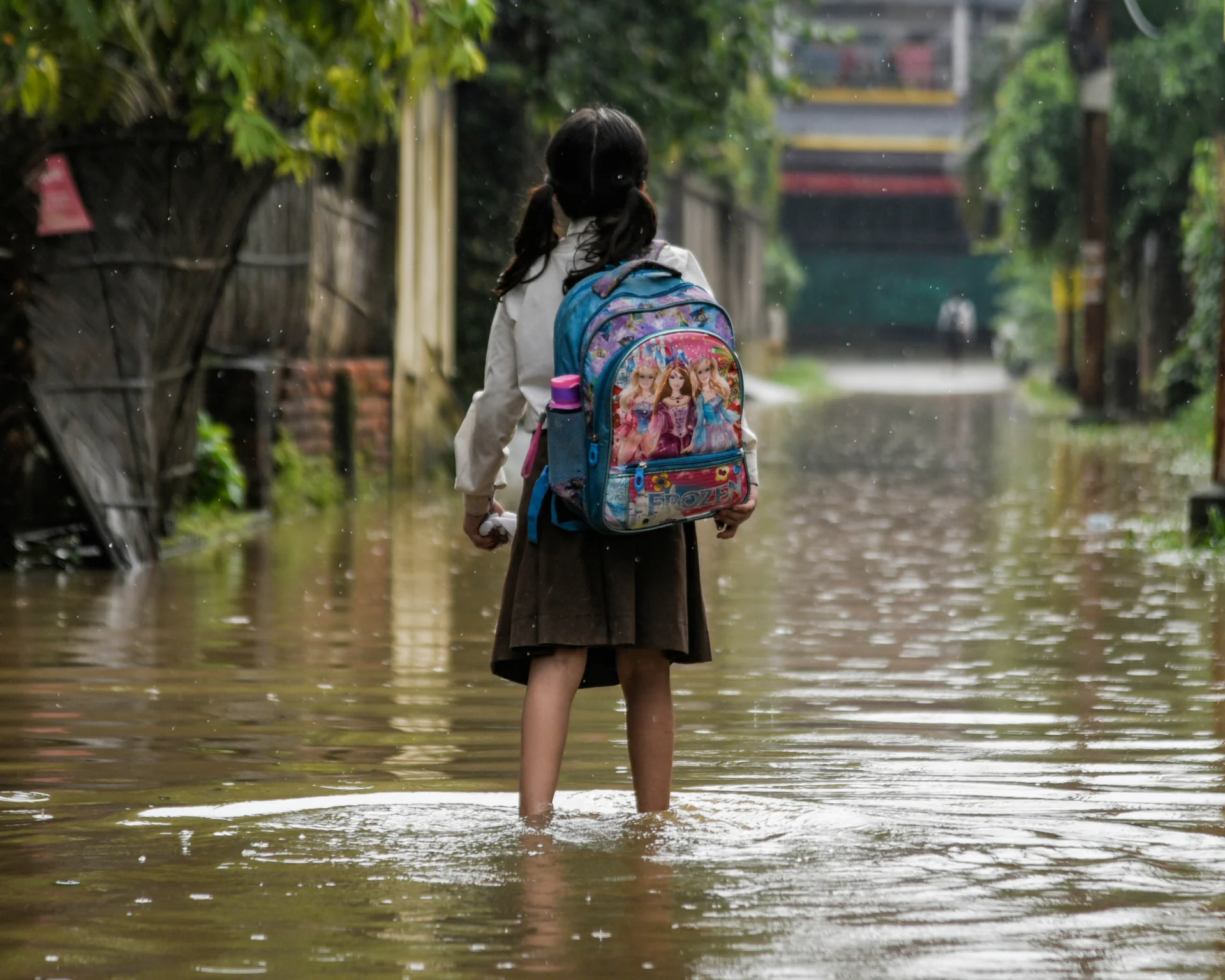
Environmental regulation could limit the amount of antibiotics contaminating soil and water by improving wastewater treatment systems, minimizing antibiotic discharge from the agriculture and pharmaceutical industries, and improving waste management. Such interventions would be particularly helpful because antibiotic resistance is most prevalent in countries where a lack of clean water and poor sanitation infrastructure accelerate the spread of antibiotic resistance. The World Health Organization reports that 1 in 3 infections in the South-East Asian and Eastern Mediterranean regions are resistant to treatment, and in Africa, 1 in 5 cases are resistant.
This risk exists because the different bacteria that affect humans, animals, and the environment do not stay in separate siloes. Waste systems, agricultural soil fertilized with manure, and rivers that collect runoff from cities and farms bring many types of bacteria into contact. In these shared environments, resistance genes can easily jump from environmental bacteria to bacteria that cause human disease. When we allow antibiotics to linger in the environment, we are maintaining a breeding ground for antibiotic resistance. Without addressing the environmental side of this crisis, antibiotic-resistant bacterial infections will continue to grow as a global health threat.